Regulation of gametophyte-to-sporophyte transitions during the life cycle
Haqin YAO
J. Mark Cock
Membres du jury:
Dr. Azeddine DRIOUICH, Rapporteur, Université de Rouen
Dr. Kenny BOGAERT, Examinateur, Université de Ghent, Belgique
Dr. Akira F. PETERS, Rapporteur, Chercheur Indépendant
Dr. J. Mark COCK, Directeur de thèse, CNRS
Dr. Susana M. COELHO, Codirectrice de thèse, CNRS
Dr. Bernad KLOAREG, Président du Jury, Sorbonne Université
Résumé/Abstract:
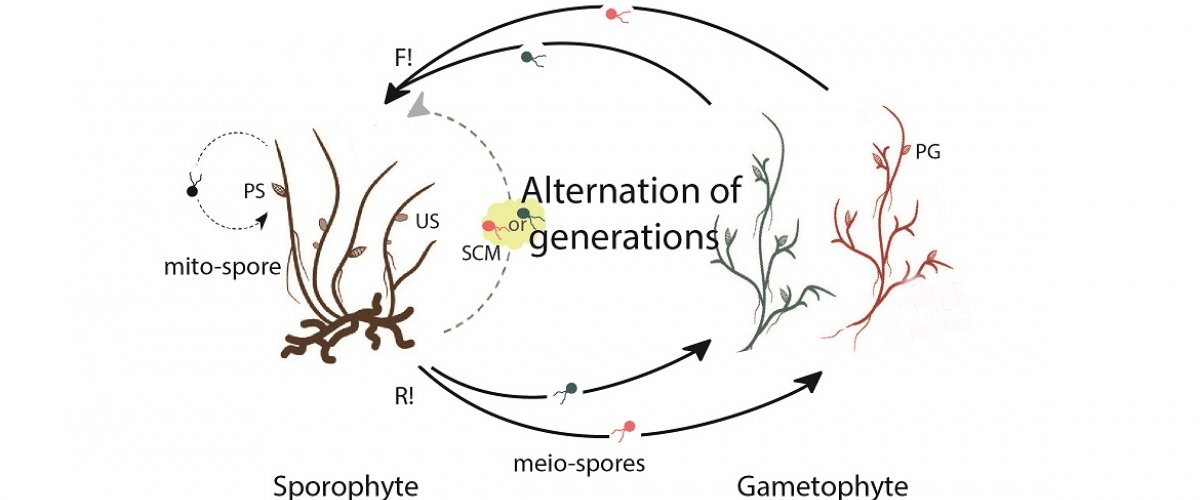
Most eukaryotic organisms reproduce sexually and have life cycles that involve an alternation between haploid and diploid phases due to two fundamental processes meiotic cell division (at the diploid-to-haploid transition) and gametes fusion or syngamy (haploid-to-diploid transition). In photosynthetic organisms with haploid-diploid life cycles, these alternations are between two distinct multicellular generations: gametophyte and sporophyte. As both the gametophyte and sporophyte generations are constructed using information from a shared genome, it follows that epigenetic regulation processes must operate both during meiosis and during syngamy to trigger the initiation of the appropriate developmental program associated with each generation. Genetic analysis of life cycle alternation in organisms diversely distribute across the lineages of the eukaryotic tree will improve our understanding at the molecular level. Current knowledge indicates that life cycle alternation is regulated by genetic factors (homeodomain transcription factors) and by chromatin modifications. The majority of brown algae have haploid-diploid life cycle and one of these species, the filamentous brown alga Ectocarpus, is being used as a model system to study life cycle regulation. Ectocarpus has a complex life cycle.
Current work has shown that alternation of generations in Ectocarpus is controlled by two homeodomain transcription factors, ORO and SAM, which regulate the induction of the sporophyte developmental program. However, alternation between the gametophyte and the sporophyte can alsobe regulated by a non-cell autonomous, sporophyte-inducing factor secreted into the culture media by sporophytes. This diffusible factor causes major developmental reprogramming in initial cells (meio- spores) of the gametophyte. Interestingly, current work shows that ORO and SAM may be part of the regulatory network triggered by the sporophyte-inducing factor. However, the biochemical nature of this factor is not known.
The main objective of this thesis was to characterize the diffusible sporophyte-inducing factor. The work focused on optimizing production, storage and bioassay of the factor and on obtaining information about its biochemical nature. The study also investigated the relationship between the sporophyte-inducing factor and two genetic regulators, ORO and SAM, to understand the developmental pathway triggered by the factor. In addition to this work on life cycle generation identity, the thesis involved characterisation of the baseless mutant, which exhibits a similar phenotype to the distag mutant and is affected in developmental patterning during both the gametophyte and sporophyte generations.