François Thomas
Chargé(e) de recherche CNRS

Marine bacteria with a sweet tooth
Microorganisms have colonized virtually all environments on Earth, and feature an incredibly diverse array of metabolic capabilities. Due to this extreme versatility, they appear as key players of biogeochemical processes, controlling fluxes of organic matter and energy in their respective ecosystems. To fully understand this contribution to the oceanic carbon cycle, we need to consider all substrates potentially available for marine microorganisms. However, unlike for small organic molecules, there is a knowledge gap on the microorganisms that can use the large, highly diverse, often complex pool of marine polysaccharides and the metabolic pathways those microbes utilize.
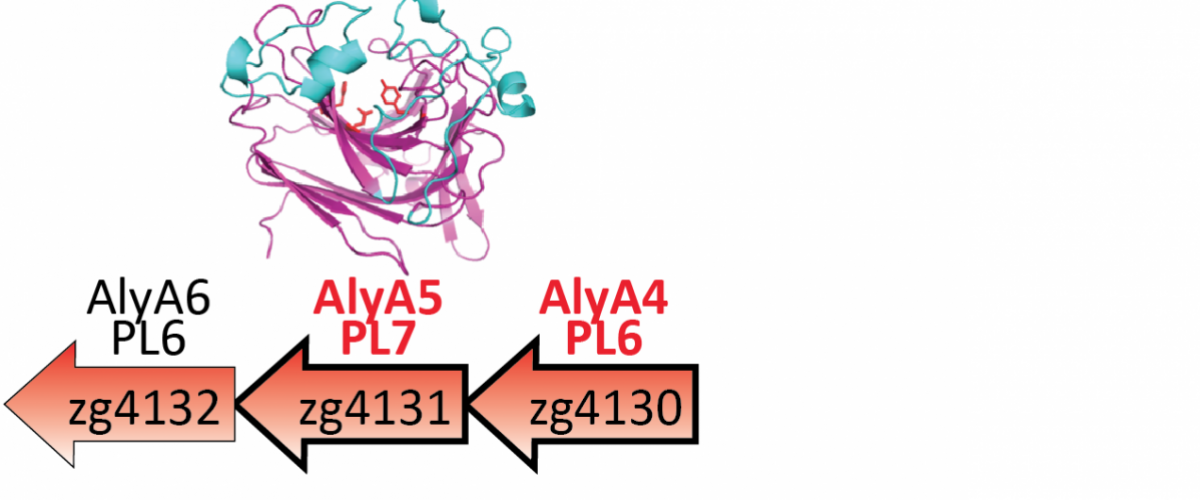
My research focuses on bacteria interacting with macroalgae, one of the main sources of polysaccharides in coastal regions worldwide. Using a combination of approaches, I investigate the identity and distribution of bacteria involved in the degradation of macroalgal polysaccharides, their specific enzymes, as well as the utilized metabolic pathways and their regulation. This strategy has already led to the discovery and biochemical characterization of the first marine operons responsible for alginate degradation, and revealed that they had surprisingly been transferred to terrestrial and human gut microbes.
Specifically, my research questions are:
-
Who are the polysaccharide-degrading bacteria (PDBac) in marine environments? Are there specialist taxa for the initial attack of HMW polysaccharides and opportunistic taxa taking up smaller degradation products?
-
How do changing habitats, spatial heterogeneities or temporal variations in polysaccharide availability shape the community assemblage and influence the activity of PDBac?
-
What are the genes involved in polysaccharide degradation and how do they evolve? Are they shared between microorganisms performing similar functions in contrasted niches?
-
What proteins do these genes encode? What are their biochemical functions in the metabolic process?
-
What are the regulatory events, controlling the expression of these genes and/or the activity of the proteins, to fine-tune the microbial response to the available substrates?
This research framework can help deciphering the yet largely unknown catabolic processes of algal polysaccharides by marine bacteria. By revealing microorganisms and pathways involved in polysaccharide degradation, it will further our fundamental understanding of organic matter recycling in marine environments, with expected discovery of new metabolic pathways and regulation strategies.
PEOPLE
Current people:
Marília Lima da Conceiçao (post-doc)
Pauline Rader (PdD student)
Kirian Gentile (M1 intern)
Former people:
Maéva Brunet (Master 2 + PhD student)
Maël Garnier (Bachelor)
Nolwen Le Duff (assistant engineer)
Manon Choulot (Master 2 student)
Jacky Ame (IUT student)
Manon Thiébaud (IUT student)
Marion Urvoy (Master 2 student)
Magda Dudek (postdoctoral scientist)
Anissa Dieudonné (Master 2 student)
Niels Zevenbergen (Master 2 student)
last update: 04/20/2023
Publications
Brott S, Nam KH, Thomas F, Dutschei T, Reisky L, Behrens M, Grimm HC, Michel G, Schweder T and Bornscheuer UT (2023) Unique alcohol dehydrogenase involed in algal sugar utilization by marine bacteria. Applied Microbiology and Biotechnology, 107, 2363-2384.
Brott S, Thomas F, Behrens M, Methling K, Bartosik D, Dutschei T, Lalk M, Michel G, Schweder T & Bornscheuer U (2022) Connecting algal polysaccharide degradation to formaldehyde detoxification. ChemBioChem 23. doi: 10.1002/cbic.202200269
Brunet M, Le Duff N, Barbeyron T & Thomas F (2022) Consuming fresh macroalgae induces specific catabolic pathways, stress reactions and Type IX secretion in marine flavobacterial pioneer degraders. The ISME Journal 16: 2027–2039. doi: 10.1038/s41396-022-01251-6
Brunet M, Le Duff N, Fuchs BM, Amann R, Barbeyron T & Thomas F (2021) Specific detection and quantification of the marine flavobacterial genus Zobellia on macroalgae using novel qPCR and CARD-FISH assays. Systematic and Applied Microbiology 44 (6): 126269. doi: 10.1016/j.syapm.2021.126269
Grigorian E, Groisillier A, Thomas F, Leblanc C & Delage L (2021) Functional characterization of a L-2-haloacid dehalogenase from Zobellia galactanivorans DsijT suggests a role in haloacetic acid catabolism and a wide distribution in marine environments. Frontiers in Microbiology 12: 725997. doi: 10.3389/fmicb.2021.725997
Barbeyron T, Thiébaud M, Le Duff N, Martin M, Corre E, Tanguy G, Vandenbol M & Thomas F (2021) Zobellia roscoffensis sp. nov. and Zobellia nedashkovskayae sp. nov., two flavobacteria from the epiphytic microbiota of the brown alga Ascophyllum nodosum, and emended description of the genus Zobellia. International Journal of Systematic and Evolutionary Microbiology, 71: 004913.
Jouanneau D, Klau LJ, Larocque R, Jaffrennou A, Duval G, Le Duff N, Roret T, Jeudy A, Aachmann FL, Czjzek M & Thomas F (2021) Structure-function analysis of a new PL17 oligoalginate lyase from the marine bacterium Zobellia galactanivorans DsijT. Glycobiology, doi: 10.1093/glycob/cwab058.
Thomas F, Le Duff N, Wu TD, Cébron A, Uroz S, Riera P, Leroux C, Tanguy G, Legeay E, Guerquin-Kern JL (2021). Isotopic tracing reveals single-cell assimilation of a macroalgal polysaccharide by a few marine Flavobacteria and Gammaproteobacteria. The ISME Journal, doi: 10.1038/s41396-021-00987-x.
Brunet M, de Bettignies F, Le Duff N, Tanguy G, Davoult D, Leblanc C, Gobet A & Thomas F (2021) Accumulation of detached kelp biomass in a subtidal temperate coastal ecosystem induces succession of epiphytic and sediment bacterial communities. Environmental Microbiology doi:10.1111/1462-2920.15389
Dudek M, Dieudonné A, Jouanneau D, Rochat T, Michel G, Sarels B & Thomas F (2020). Regulation of alginate catabolism involves a GntR family repressor in the marine flavobacterium Zobellia galactanivorans DsijT. Nucleic Acids Research 48 (14): 7786-7800.
de Bettignies F, Dauby P, Thomas F, Gobet A, Delage L, Bohner A, Loisel S & Davoult D (2020) Degradation dynamics and processes associated with the accumulation of Laminaria hyperborea (Phaeophyceae) kelp fragments: an in situ experimental approach. Journal of Phycology DOI: 10.1111/jpy.13041-19-156
Thomas F, Le Duff N, Leroux C, Dartevelle L & Riera P (2020). Isotopic labeling of cultured macroalgae and isolation of 13C-labeled cell wall polysaccharides for trophic investigations. Advances in Botanical Research, 55: 1-17.
Thomas F, Dittami S, Brunet M, Le Duff N, Tanguy G, Leblanc C & Gobet A (2019) Evaluation of a new primer combination to minimize plastid contamination in 16S rDNA metabarcoding analyses of alga‐associated bacterial communities. Environmental Microbiology Reports, doi:10.1111/1758-2229.12806
Thomas F, Morris JT, Wigand C & Sievert SM (2019) Short-term effect of simulated salt marsh restoration by sand-amendment on sediment bacterial communities. Plos One 14(4) e0215767
Thomas F, Corre E & Cébron A (2019) Stable isotope probing and metagenomics highlight the effect of plants on uncultured phenanthrene-degrading bacterial consortium in polluted soil. The ISME Journal, doi: 10.1038/s41396-019-0394-z.
McNichol J, Stryhanyuk H, Sylva SP, Thomas F, Musat N, Seewald JS & Sievert SM (2018) Primary productivity below the seafloor at deep-sea hot springs. PNAS, doi: 10.1073/pnas.1804351115.
Ficko-Blean E*, Préchoux A*, Thomas F, Rochat T, Larocque R, Zhu Y, Stam M, Génicot S, Jam M, Calteau A, Viart B, Ropartz D, Pérez-Pascual D, Correc G, Matard-Mann M, Stubbs KA, Rogniaux H, Jeudy A, Barbeyron T, Médigue C, Czjzek M, Vallenet D, McBride MJ, Duchaud E & Michel G (2017) Carrageenan catabolism is encoded by a complex regulon in marine heterotrophic bacteria. Nature Communications, doi:10.1038/s41467-017-01832-6. *Equal contributions
Thomas F, Bordron P, Eveillard D & Michel G (2017) Gene expression analysis of Zobellia galactanivorans during the degradation of algal polysaccharides reveals both substrate-specific and shared transcriptome-wide responses. Frontiers in Microbiology, 8:1808. doi: 10.3389/fmicb.2017.01808
Zhu Y*, Thomas F*, Larocque R, Li N, Duffieux D, Cladière L, Souchaud F, Michel G & McBride MJ (2017) Genetic analyses unravel the crucial role of a horizontally acquired alginate lyase for brown algal biomass degradation by Zobellia galactanivorans. Environmental Microbiology, 19:6, 2164-2181. doi: 10.1111/1462-2920.13699. *Equal contributions
Ritter A, Cabioch L, Brillet-Guégen L, Corre E, Cosse A, Dartevelle L, Duruflé H, Fasshauer C, Goulitquer S, Thomas F, Correa JA, Potin P, Faugeron S & Leblanc C (2017) Herbivore-induced chemical and molecular responses of the kelps Laminaria digitata and Lessonia spicata. PLoS one, 12:3.
Bourceret A, Leyval C, Thomas F & Cébron A (2017) Rhizosphere effect is stronger than PAH concentration on shaping spatial bacterial assemblages along centimeter-scale depth gradients. Canadian Journal of Microbiology, doi: 10.1139/cjm-2017-0124
Barbeyron T, Thomas F, Barbe V, Teeling H, Schenowitz C, Dossat C, Goesmann A, Leblanc C, Glöckner FO, Czjzek M, Amann R & Michel G (2016) Habitat and taxon as driving forces of carbohydrate catabolism in marine heterotrophic bacteria: example of the model algae-associated bacterium Zobellia galactanivorans DsijT. Environmental Microbiology, 18:12, 4610-4627.
Thomas F & Cébron A (2016) Short-term rhizosphere effect on available carbon sources, phenanthrene degradation, and active microbiome in an aged-contaminated industrial soil. Frontiers in Microbiology, 7:92.
Thomas F, Lorgeoux C, Faure P, Billet D & Cébron A (2016) Isolation and substrate screening of polycyclic aromatic hydrocarbon degrading bacteria from soil with long history of contamination. International Biodeterioration and Biodegradation, 107, 1-9.
McNichol J, Sylva SP, Thomas F, Taylor CD, Sievert SM & Seewald JS (2016) Assessing microbial processes in deep-sea hydrothermal systems by incubation at in situ temperature and pressure. Deep Sea Research Part I, 115, 221-232.
Signori CN*, Thomas F*, Prast AE, Pollery RCG & Sievert SM (2014) Microbial diversity and community structure across environmental gradients in Bransfield Strait, Western Antarctic Peninsula. Frontiers in Microbiology, 5:647. *Equal contributions
Thomas F, Giblin AE, Cardon ZG & Sievert SM (2014) Rhizosphere heterogeneity shapes abundance and activity of sulfur- oxidizing bacteria in vegetated salt marsh sediments. Frontiers in Microbiology, 5:309.
Thomas F, Cosse A, Le Panse S, Kloareg B, Potin P & Leblanc C (2014) Kelps feature systemic defense responses: insights into the evolution of innate immunity in multicellular eukaryotes. New Phytologist, 204:3, 567-576.
Thomas F, Lundqvist LCE, Jam M, Barbeyron T, Sandström C, Michel G & Czjzek M (2013) Comparative characterization of two marine alginate lyases from Zobellia galactanivorans reveals distinct modes of action and exquisite adaptation to their natural substrate. The Journal of Biological Chemistry, 288:32, 23021-37.
Thomas F, Barbeyron T, Tonon T, Czjzek M & Michel G (2012) Characterization of the first alginolytic operons in a marine bacterium: From their emergence in marine Flavobacteriia to their independent transfers to marine Proteobacteria and human gut Bacteroides. Environmental Microbiology, 14:9, 2379-94.
Hehemann JH, Correc G, Thomas F, Bernard T, Barbeyron T, Jam M, Helbert W, Michel G & Czjzek M (2012) Biochemical and structural characterization of the complex agarolytic system from the marine bacterium Zobellia galactanivorans. The Journal of Biological Chemistry, 287:36, 30571-84.
Thomas F, Hehemann JH, Rebuffet E, Czjzek M & Michel G (2011) Environmental and gut Bacteroidetes: the food connection. Frontiers in Microbiology, 2:93.
Thomas F, Barbeyron T & Michel G (2011) Evaluation of reference genes for real time quantitative PCR in the marine flavobacterium Zobellia galactanivorans. Journal of Microbiological Methods, 84, 61-66.
Thomas F, Cosse A, Goulitquer S, Raimund S, Morin P, Valero M, Leblanc C & Potin P (2011) Waterborne signaling primes the expression of elicitor-induced genes and buffers the oxidative responses in the brown alga Laminaria digitata. PLoS one, 6:6.
McFiggans G, Bale CSE, Ball SM, Beames JM, Bloss WJ, Carpenter LJ, Dorsey J, Dunk R, Flynn MJ, Furneaux KL, Gallagher MW, Heard DE, Hollingsworth AM, Hornsby K, Ingham T, Jones CE, Jones RL, Kramer LJ, Langridge JM, Leblanc C, LeCrane JP, Lee JD, Leigh RJ, Longley I, Mahajan AS, Monks PS, Oetjen H, Orr-Ewing AJ, Plane JMC, Potin P, Shillings AJL, Thomas F, von Glasow R, Wada R, Whalley LK & Whitehead JD (2010). Iodine-mediated coastal particle formation: an overview of the Reactive Halogens in the Marine Boundary Layer (RHaMBLe) Roscoff coastal study. Atmospheric Chemistry and Physics, 10, 2975–2999.
Goulitquer S, Ritter A, Thomas F, Ferec C, Salaün JP & Potin P (2009) Release of volatile aldehydes by the brown algal kelp Laminaria digitata in response to both biotic and abiotic stress. Chembiochem, 10, 977-982.
Alfaro AC, Dewas SE & Thomas F (2007) Food and habitat partitioning in grazing snails (Turbo smaragdus), northern New Zealand. Estuaries and Coasts, 30, 431-440.
Alfaro AC, Thomas F, Sergent L & Duxbury M (2006) Identification of trophic interactions within an estuarine food web (northern New Zealand) using fatty acid biomarkers and stable isotopes. Estuarine and Coastal Shelf Science, 70, 271-286.